“Jumping genes” are ubiquitous. Every domain of life hosts these sequences of DNA that can “jump” from one position to another along a chromosome; in fact, nearly half the human genome is made up of jumping genes. Depending on their specific excision and insertion points, jumping genes can interrupt or trigger gene expression, driving genetic mutation and contributing to cell diversification. Since their discovery in the 1940s, researchers have been able to study the behavior of these jumping genes, generally known as transposons or transposable elements (TE), primarily through indirect methods that infer individual activity from bulk results. However, such techniques are not sensitive enough to determine precisely how or why the transposons jump, and what factors trigger their activity.
Reporting in the Proceedings of the National Academy of Sciences, scientists at the University of Illinois at Urbana-Champaign have observed jumping gene activity in real time within living cells. The study is the collaborative effort of physics professors Thomas Kuhlman and Nigel Goldenfeld, at the Center for the Physics of Living Cells, a National Science Foundation Physics Frontiers Center.
“These are genes that hop around and change location within the genome of a cell,” said Kuhlman. “We hooked that activity up to a molecular system, such that when they start hopping around, the whole cell fluoresces. In our experiment, cells fluoresced most when they weren’t very happy. One school of thought suggests that an increased mutation rate like this might be an advantage in such unhappy conditions, for cells to diversify.”
In order to help design the experiment and to calculate what would have happened if the jumping occurred in a purely random fashion, Goldenfeld’s team developed computer simulations of the growth of bacterial colonies and predicted what the experimental signal would look like in the random case. These calculations showed that the experiments could not simply be interpreted as random transposon activity and even provided clues as to sources of non-randomness, including environmental feedback and heredity.
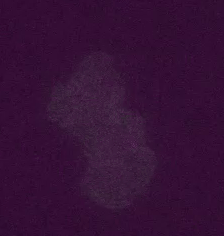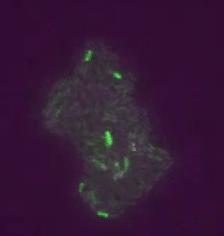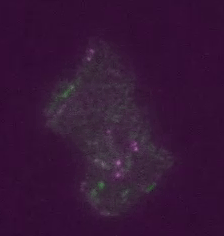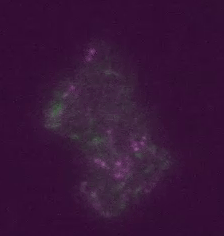
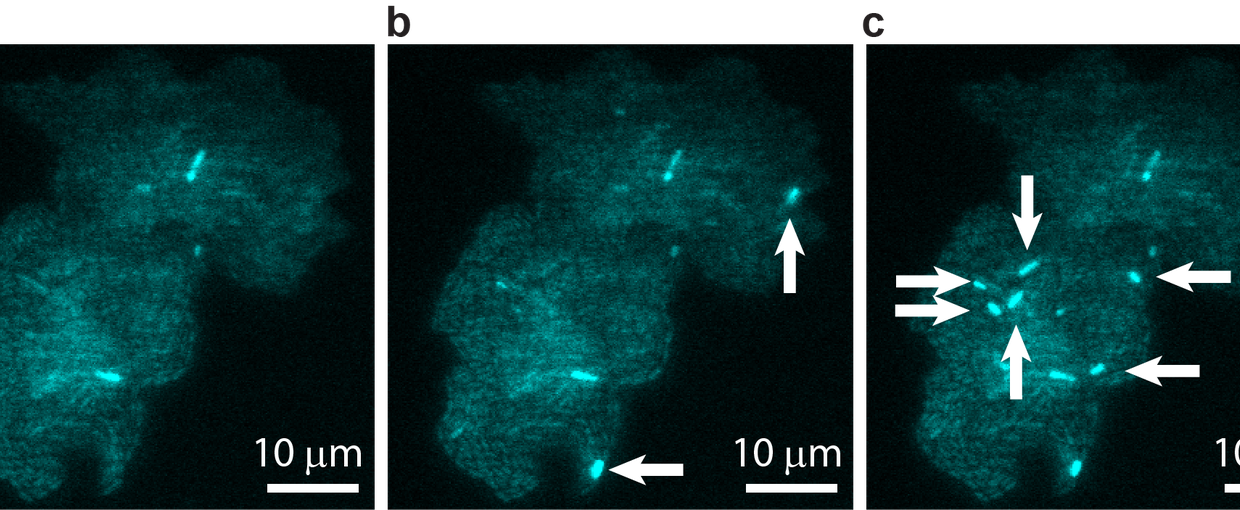
Real time transposable element activity in a small colony of bacteria, where excision of the transposable element causes the cell to light up (false colored green), and the excision reaction is caused by transposase protein (false colored magenta). Video courtesy of T.E. Kuhlman, University of Illinois at Urbana-Champaign, reproduced with permission from Proceedings of the National Academy of Sciences USA.
“Our work involved a good deal of computational image analysis, followed by statistical analysis,” said Goldenfeld. “To extract signals and conclusions from the raw data, simulation and theoretical calculations were integral to the experimental design and interpretation. This sort of collaborative project was really only possible with the unique structure provided by the Center for the Physics of Living Cells.”
“The over-arching long-term research goal is a deeper understanding of how evolution works at the molecular level. Direct observation of how genomes in cells restructure themselves allows for a precise determination of adaptation rates and may shed light on a host of important evolutionary questions, ranging from the emergence of life to the spread of cancer, where cells undergo rapid mutations and transformations of their genomes,” added Kuhlman.
E. coli cells carrying a “wildtype” version of the transposable element grow in a monolayer within a microfluidic device; a single cell is seen to light up green upon experiencing a transposable element event. Video courtesy of T.E. Kuhlman, University of Illinois at Urbana-Champaign.
Kuhlman and Goldenfeld are members of the Carl R. Woese Institute for Genomic Biology and the Department of Physics at the University of Illinois at Urbana-Champaign. The work involved several graduate students, including lead author Hyuneil Kim, Gloria Lee, Nicholas Sherer and Michael Martini.
This release is based upon research performed at the Center for the Physics of Living Cells, a Physics Frontiers Center supported by the National Science Foundation, and at the NASA Astrobiology Institute for Universal Biology supported by NASA, both located on the University of Illinois at Urbana-Champaign campus. Kuhlman’s research is additionally supported by a 2015 Sloan Research Fellowship from the Alfred P. Sloan Foundation. The conclusions presented are those of the scientists, and not necessarily those of the funding agencies.