
May 24, 2016
Program News
New Insight into Microbial Communities in Anoxic Sediments
Metagenomic analyses of an autotrophic Fe(II)-oxidizing, nitrate-reducing enrichment culture.
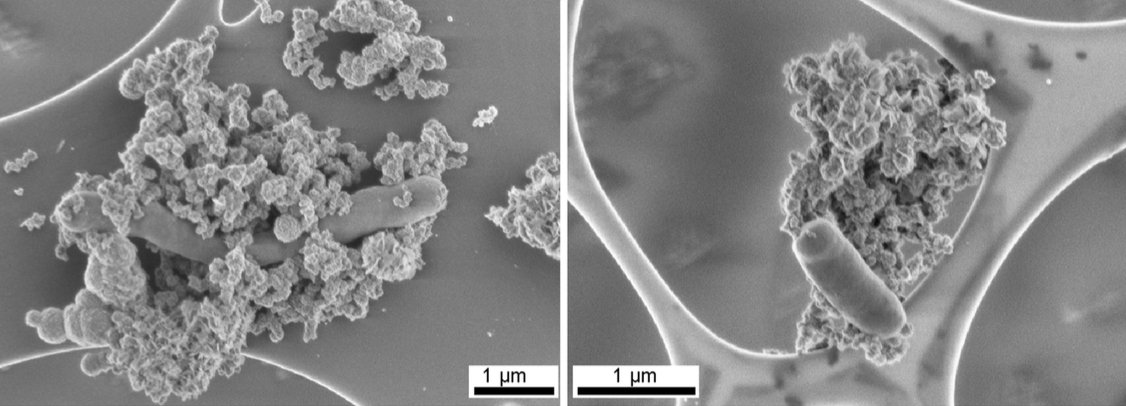

Scanning electron micrographs of anaerobic Fe(II)-oxidizing cultures. From: Schadler et al. (2009) Formation of Cell-Iron-Mineral Aggregates by Phototrophic and Nitrate-Reducing Anaerobic Fe(II)-Oxidizing Bacteria, Geomicrobiology Journal. Image credit: Reprinted by permission of Taylor & Francis LLC, (http://www.tandfonline.com).
A new study provides insight into a well-recognized chemolithotrophic pathway that can be used by microorganisms inhabiting anoxic sediments. Researchers used an enrichment culture of chemolithoautotrophic organisms from freshwater sediments (dubbed Culture KS) as a model system to study the pathway: nitrate-dependent ferrous iron [Fe(II)] oxidation (NDFO).
Previous efforts to isolate the organism responsible for the oxidation of iron [Fe(II)] in Culture KS had proven unsuccessful. In the new study, researchers used metagenomic analysis to better understand the roles of different bacteria in Culture KS. Using this method, they were able to identify the primary Fe(II)-oxidizer as a species in the family Gallionellaceae and to obtain a near-complete genome from the organism. In addition, draft genomes from other community members were acquired.
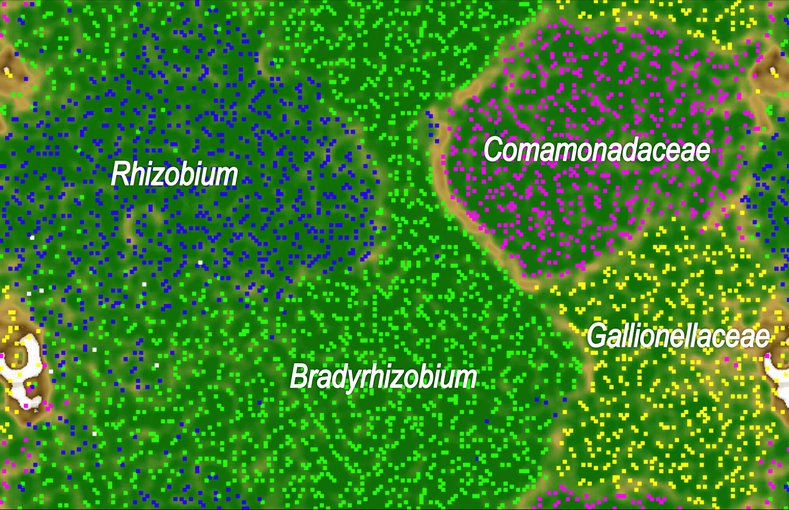
An emergent self-organizing map (ESOM) calculated using tetranucleotide frequencies of contigs in the metagenome.Image credit: He et al. (2016).
The team then studied these genomes and identified genes involved in electron transfer pathways. In doing so, they were able to map out interactions between different species of microorganisms in Culture KS. The results show that the metabolisms of different species in the culture are interdependent, and these organisms cooperate to successfully achieve NDFO.
The iron oxidation mechanisms and microbial interactions revealed by this study help to inform our understanding of iron-based chemolithoautotrophic microbial pathways on Earth and how they might be operative on other iron-rich rocky planets.
The paper, “Metagenomic analyses of the autotrophic Fe(II)-oxidizing, nitrate-reducing enrichment Culture KS,” was published in the journal Applied Environmental Microbiology. The research was supported by the NASA Astrobiology Institute element of the NASA Astrobiology Program.