Written byMichael Schirber

Languages aren’t set in stone. They constantly mutate, adding new words and modifying old meanings. And they also diversify into multiple dialects, with their own idioms and pronunciations. Some of these dialects diverge so far from the mother tongue that they become their own separate language.
This evolution in languages has a lot in common with evolution in organisms. Eric Smith, a complex systems researcher at the Santa Fe Institute, studies both language and biology. He believes there are lessons from linguistics that may help biologists in their efforts to map out the history of life on Earth.
“In both linguistics and biology, people are reasoning about the past,” Smith says. “They are creating narratives to explain why there is so much diversity in languages or in organisms.”
In trying to understand languages, linguists look for commonalities and differences between languages that may hint at relationships in the past. This information can be collected into an evolutionary tree, similar to what is done in biology.
However, the tools for building trees are very different in the two disciplines. Modern biologists use highly sophisticated computational methods to relate DNA sequences in the genomes of different organisms. This focus on DNA has given biologists a kind of tunnel vision, Smith says. They see the trees but miss the forest.
By contrast, linguists take a more systematic approach, looking at language in all its facets — not simply focusing on some grammatical code or pronunciation key.
“It is very hard to make the same oversimplifications for languages,” Smith says. “One generally needs to be thinking about the role of all linguistic elements: lexical roots, sounds, syntax, etc.”
Smith—who is supported by funding from NASA’s Astrobiology Institute—has been working on some of these linguistic elements, trying to build evolutionary trees from the information they contain. He believes this work may provide biologists with insights into how they can broaden their perspective to include more than DNA alphabets in their narratives of life’s past.

Eric SmithImage credit: Eric Smith.
Language under the microscope
Linguists build evolutionary trees for the same reason that biologists do: The roots and branches hold information about the past. In the case of language, the tree shape can tell us about migrations, exchanges and cultural divides.
When linguists construct a tree, their starting point is basically the same as in biology.
“You look at several copies of essentially the same thing and then try to account for the variability,” Smith explains.
In language, the objects of investigation are often sound units (or phonemes) that make up words. Phonemes are in some ways like the nucleotide bases (A,T,C,G) of DNA. Pronouncing a string of phonemes together gives a word, just as a string of bases defines a gene. And just as mutations in a gene sequence can lead to the emergence of a new species, changes in phoneme usage can result in the formation of new languages.
Take the different words for the number two. In English, we say “two,” but in German it’s “zwei,” in French “deux,” and in Spanish “dos.” Linguists have found that vowels often evolve more rapidly than consonants, so one can infer that the emerging languages of French and Spanish probably had the same word for two in the relatively recent past. By contrast, the distinction between the German/English “zw/t” and the French/Spanish “d” implies that the ancestors that gave rise to their language families split from each other at a much earlier time.
Similar types of inferences are made in genomics, where some types of DNA mutations are considered more probable than others. However, there is an important difference between biological evolution and language evolution. Genetic changes can occur in isolation, with a single flip in one DNA letter while the rest of the genome stays the same. This independent, or “sporadic,” evolution is highly unlikely in language. A particular sound is identified by the collection of words in which it is pronounced, so when that sound changes, it typically changes in many words at the same time.
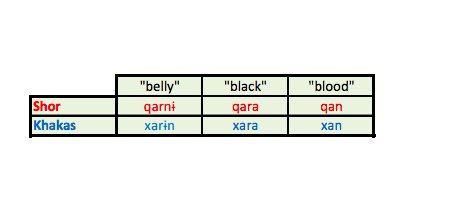
Examples of words from the Turkic language Shor where the letter q has become a letter x in the related language of Khakas.Image credit: derived from Current Biology 25, 1–9, January 5, 2015.
Concerted effort
The collective behavior found in language evolution has its counterpart in biology. One can find instances where several genes appear to have evolved in unison. This so-called “concerted” evolution occurs when relationships within a genome prevents individual genes from mutating without parallel changes in other genes.
It is not always clear when concerted evolution applies, rather than the more common sporadic evolution. Two similar-looking genetic sequences could signify some sort of mutual history, or they could just be the result of random chance.
To explore this issue, Smith and his colleagues considered sporadic versus concerted evolution in a language context, where it’s far clearer the role that system-wide connections play. As reported in a 2015 Current Biology paper, the team analyzed 62 distinct speech sounds in 26 Turkic languages spoken across Asia, identifying numerous sound changes in a wide variety of words.
As an example, the team observed 34 instances where a word in the Siberian language Shor has a “q” whereas the same word in the neighboring language of Khakas has an “x.” If this had been genomics instead of language, one could quite naturally assume that these 34 changes in word pronunciations occurred independently. However, Smith and his collaborators showed that this sporadic evolution model produces a faulty tree. Specifically, it predicts that these Turkic languages originated around 2,400 BCE, which is 2,000 years older than other historical estimates.
The researchers argued that a better starting assumption is the one that makes most sense linguistically: that sound changes occur in conjunction, rather than independently. The early Khakas speakers switched “q” to “x” in several words at roughly the same time. Smith and his coauthors devised a method—based on Markov chain Monte Carlo statistics—for identifying such concerted change events in genetic or linguistic data. They demonstrated their method with the Turkic languages, creating a language tree that was consistent with other cultural information.
More recently, Smith and co-workers performed a linguistic analysis of a wide range of languages based on word meanings. As described in a 2016 Proceedings of the National Academy of Sciences paper, they found similarities in how different languages group words together based on meaning, but they also found that these groups could shift over time. Such meaning-shifts might be used in the future to build language trees, as is done now with sound-changes.
An evolution in evolutionary studies
In the bigger picture, Smith believes that language studies can show biologists how to go beyond the current paradigm of “Evolution 1.0”.
The standard method for seeing into the genetic past is called “phylogenetic reconstruction.” It involves taking similar copies of the same gene in two related organisms and then to imagine what DNA-altering steps, such as mutations and transpositions, might have led to these genes diverging from each other. There are many possibilities, so biologists assign probabilities to each step and compute the most likely path. The end result is an estimate of when the two organisms diverged from each other and how the given gene looked in their common ancestor.
The nice thing about phylogenetic reconstruction is it gives a concrete, reproducible answer. But Smith thinks this should only be considered a first draft. The probability assignments can provide guidance, but they are only a crude tool. “It is very hard to make models complex enough to capture things we believe happen in the world,” Smith says.

Greg Fournier of MIT.Image credit: MIT.
Greg Fournier a biologist from MIT believes Smith is right in his assessment of Evolution 1.0. Fournier, who works on genetic reconstructions of microbial lineages, says one of the common over-simplifications is the principle of parsimony, which basically instructs biologists to choose the evolutionary path with the least number of changes. “But evolution is neutral,” Fournier says. “Life can evolve over an enormous realm of possibilities, so it likely takes a random walk, rather than the shortest path.”
Taking the shortest path can sometimes produce nonsensical intermediate steps. For example, imagine a path from a present-day gene to an ancestor gene that goes through some intermediate gene X, but gene X produces a faulty protein. “Our reconstructions may traverse spaces that are non-functional,” Fournier says.
It is much harder to slip into this kind of mistake in linguistics, Smith explains, because linguists are constantly aware that each evolutionary step must give a working language.
“There are many constraints on how languages may or may not actually be put together, and many more constraints on how they may or may not change,” Smith says.
Biology could incorporate similar sorts of constraints, as Smith demonstrated with Rogier Braakman, another biologist from MIT, in a study of carbon fixation pathways. These pathways are chemical reactions used by organisms to convert carbon dioxide into organic molecules for use in cells. Biologists know of six different pathways, but it’s been unclear how they relate to each other in an evolutionary sense.

Rogier Braakman of MIT.Image credit: MIT.
In their study, published in 2012 in the journal PLoS Computational Biology, Braakman and Smith built a tree with the constraint that the carbon fixation function has been a continuous feature of life on Earth. In other words, species come and go, but there has always been some set of organisms that can fix carbon, otherwise the ecological system would have collapsed. Using this functional constraint, Braakman and Smith showed that innovations in carbon fixation could explain many of the early branches in the tree of life.
“We found that function does play an important role in giving the outline of the tree of life,” Braakman says.
Smith says other researchers are adding similar types of constraints to their statistical models. Fournier, for example, is looking into incorporating protein-folding constraints into his work. However, Smith says that evolving from the current 1.0 version to Evolution 2.0 won’t be an easy task. One really has to stand back and think about the problem on a large scale. Making analogies with language evolution can help because the use of constraints and other systemic approaches are so clearly vital in reconstructing our linguistic past.
“Even though our language work applies to a different data domain than much of what we might do with genes, organisms, or ecosystems, many of the principles of getting back to innovating in evolutionary reasoning, rather than just repeatedly using packaged approaches that are now established, should be relevant,” Smith says.

