The eukaryotes are a sophisticated bunch. In their ranks, one finds multi-cellular organisms like animals, plants, and fungi. Even the uni-cellular eukaryotes, such as amoebas and yeast, have an intracellular complexity that far exceeds the simple machinery inside prokaryotes, classified as the “non-eukaryotes.” Biologists have been debating the origin of eukaryotic complexity for decades, but a study published last year of deep ocean deposits may have uncovered an evolutionary clue within a prokaryote.
In a sample taken from the Arctic Ocean, researchers have identified a microbial organism belonging to the domain of Archaea. What sets this prokaryote apart is that it has genetic markers that put it closer to eukaryotes than any other prokaryote studied before. Moreover, this organism, which has been named Lokiarchaea, appears to have genes that are typically associated with eukaryotic functions, such as membrane remodeling capabilities.
This finding fits into a theory that eukaryotes evolved from an archaeal ancestor, making Lokiarchaeota a kind of “missing link” in the universal tree of life.
“If it is real, the finding of Lokiarchaea may show a closer relationship between archaea and eukaryotes,” says microbiologist Rachel Whitaker of University of Illinois. “That would mean the split between eukaryotes and archaea might have been more recent than thought.”
But these claims are disputed by some scientists, who see problems in the genetic analysis of Lokiarchaeota. They contend that eukaryotes evolved from a more ancient organism at the base of the tree of life. This debate is not a new one. It’s been going on for decades but seems no closer to resolution. In fact, biologists can’t even agree on what to call the two sides of the debate. The archaeal ancestor hypothesis is often called the two-domain model, but it also goes by the name eocyte model or merger model. The opposing viewpoint is often referred to as the three-domain model, or the deep root model.
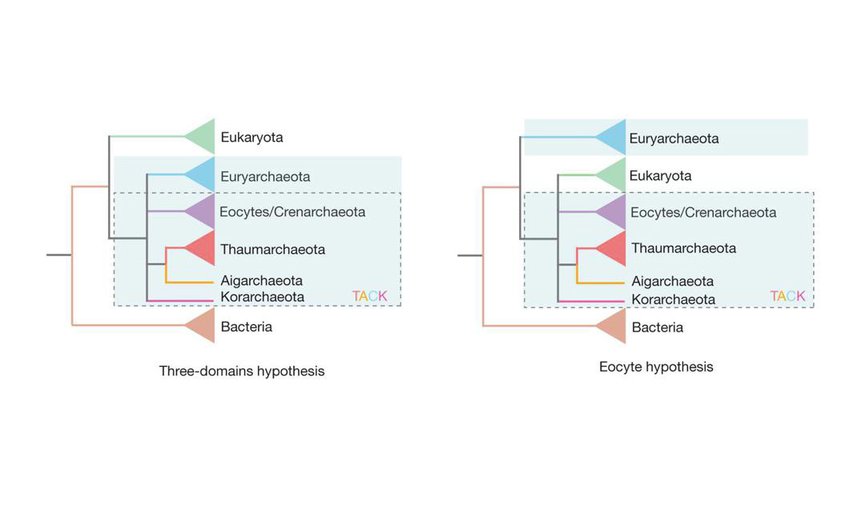
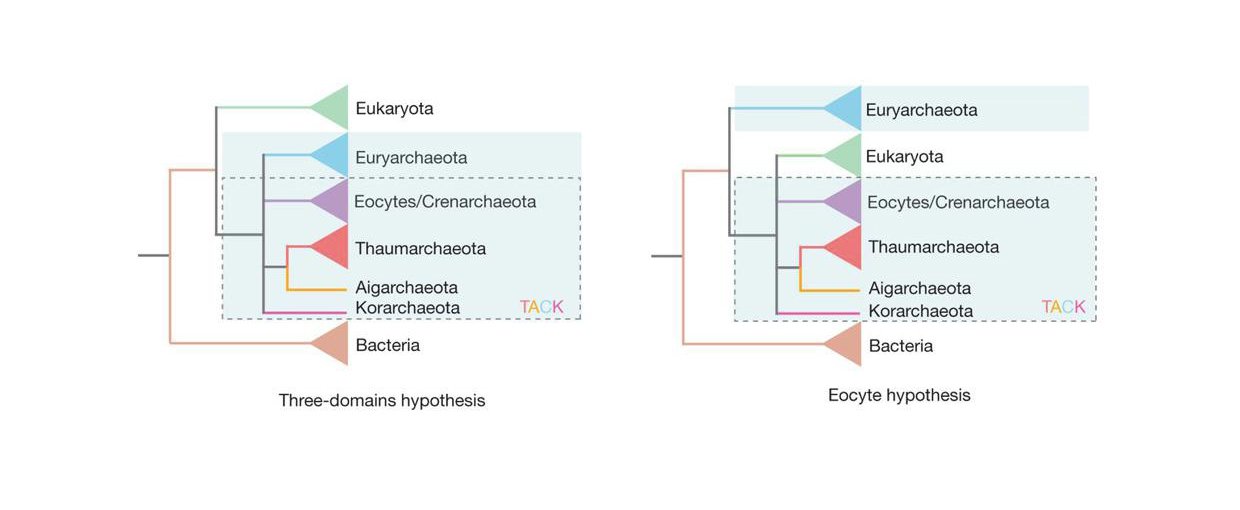
The three-domain model on the left shows Bacteria, Eukaryota, and Archaea branching close to the root of the tree of life. Note that Archaea (blue-shaded box) is divided into different groups, some of which form the TACK supergroup (dashed-line box). On the right is the two-domain, or eocyte, model. In this case, Eukaryota is one branch within Archaea.Image credit: Williams et al., Nature 504, 231–236 (2013); http://www.nature.com/nature/journal/v504/n7479/full/nature12779.html.
Name game
Naming (and renaming) things has important implications in biology as to how organisms are classified. The name eukaryote is no exception. In Greek, the word signifies “true kernel,” and is meant to draw attention to the most distinguishing characteristic of the eukaryote cell: its nucleus. This membrane-bound vesicle neatly separates the cell’s DNA from the rest of the cellular activity. Prokaryotes do not have such a compartment, and their name, which means “before kernel,” reflects this missing trait.
The eukaryote-prokaryote naming convention became dominant in the 1960s. Its usage contained a hidden bias: the pro-karyotes seem so primitive, they must have come before the eukaryotes. That bias was challenged by the late Carl Woese. In the 1970s, he was studying the 16S rRNA gene that codes for ribosomal RNA. This gene is found in all prokaryotes, where it is highly conserved (meaning it has not evolved very much). By comparing small changes in the 16S rRNA sequence, Woese and his colleagues were able to determine how organisms across the tree of life are related. From this “phylogenetic” analysis, they discovered that a subset of prokaryotes, called archaea, were as far apart from bacteria as either was distant from eukaryotes.
In light of these findings, Woese proposed in 1977 a three-domain tree of life, with Bacteria, Archaea, and Eukaryota branching out from a single common root. The primitive organism at the base, the so-called last universal common ancestor (LUCA), is thought to have lived 3.5 billion years ago.
Not long after the three-domain proposal, other scientists started to find interesting connections across domains. In 1984, the biologist James Lake from the University of California Los Angeles and his collaborators pointed out structural similarities in the ribosomes of eukaryotes and certain archaea, which were called “eocytes” or dawn cells. Subsequent analysis of certain eocyte genes showed that they were very similar to the corresponding genes in eukaryotes, suggesting a common ancestry.
“I’ve always known that eocytes were the source of the eukaryotic nuclear (informational) genes and been confident of it,” Lake says.
The eocyte findings imply that Eukaryota are not a separate domain, but rather a branch inside Archaea. Lake believes the acceptance for the eocyte hypothesis has “skyrocketed” over the past decade due to phylogenetic work with other genes besides 16S rRNA. This research has led to several different variants of the two-domain model, some of which propose that the eukaryotes originated from a cell merger between an ancient archaea and an early form of bacteria. The timing of the first eukaryote’s arrival is pegged at around 2 billion years ago (more than a billion years later than the estimate from the three-domain model).
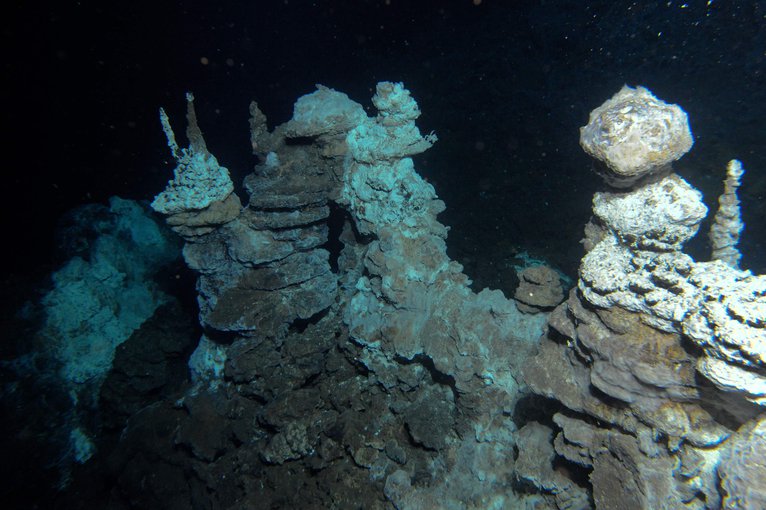

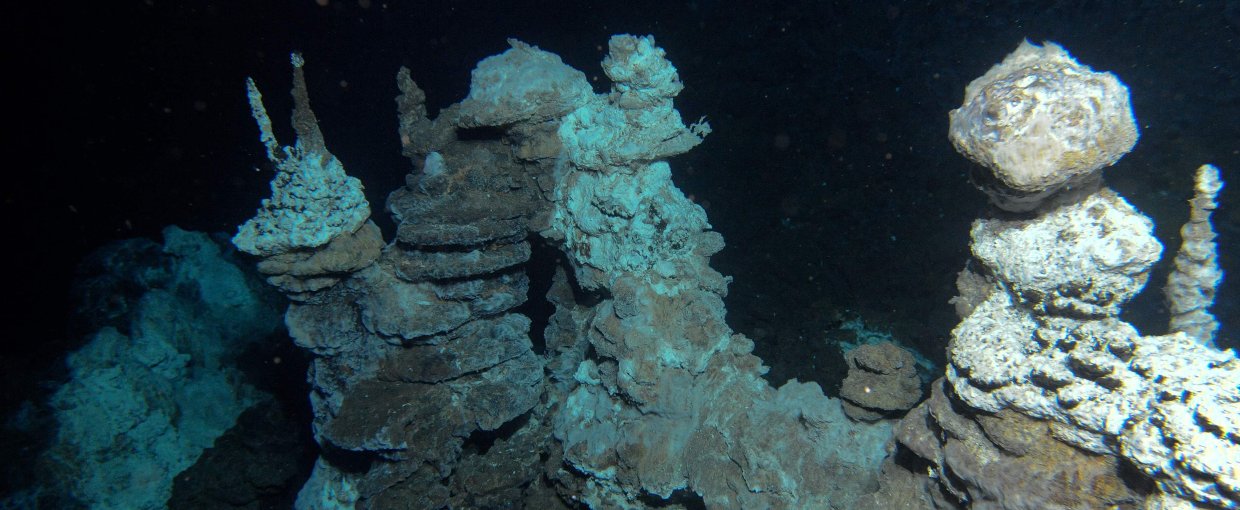
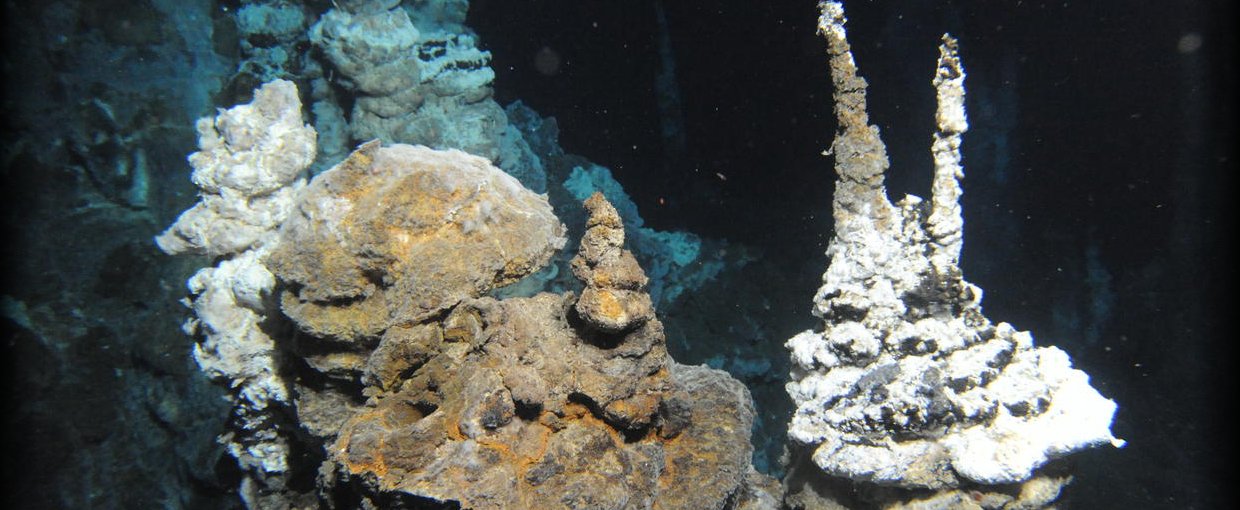
Loki’s Castle is a hydrothermal vent field along the Arctic Mid-Ocean Ridge discovered by researchers from the Centre for Geobiology in Norway. Image credit: Centre for Geobiology by R.B. Pedersen.
Microbial dark matter
For the two-domain tree of life, different models have the eukaryotes branching in different places. Currently, genetic evidence suggests that eukaryotes arose out of the TACK supergroup of Archaea. These organisms are not yet well-understood, partly because biologists have a hard time culturing them in a lab. These slow-growing archaea live under harsh conditions and therefore tend not to thrive in “captivity.”
Many other organisms are in a similar boat. They can’t be cultured, so they often go unnoticed. But many biologists are now concentrating their efforts on this so-called “microbial dark matter.” They use genetic markers to target these organisms in the wild and then sequence whatever DNA they can find to build up a genome. Here is where the story comes back to Lockiarchaeota.
A team of microbiologists, led by Thijs Ettema of Uppsala University in Sweden, were studying microbial dark matter from a subset of the TACK supergroup, called the Deep-Sea Archaeal Group (DSAG). Little is known about the DSAG, but Ettema and his colleagues hoped that these deep-ocean-dwelling organisms might offer new insights into the origin of the eukaryotic cell.
The researchers obtained a sample of marine deposits from the Arctic Mid-Ocean Spreading Ridge, at a depth of 3,000 meters. The team performed a preliminary test by sequencing 16S rRNA gene sequences and found that about 10 percent of the organisms in the sample were members of DSAG.
Ettema’s team tried to isolate a single cell for genomic sequencing, but this turned out be a “dead end,” Ettema says. As an alternative, they laid a wide net and sequenced all the DNA in the sample. This so-called metagenomic analysis is “contaminated” with the DNA of non-target organisms. “There can be hundreds or thousands of organisms in a sample, so it might seem like ‘mission impossible’ to reconstruct a single genome,” Ettema says.
However, over the years, microbiologists have developed tools to separate the apples from oranges in the sequencing data. One of these tools is metagenomic binning, which uses frequencies of DNA snippets of defined length (typically 4 bases long) to group together metagenomic DNA fragments of the same species.
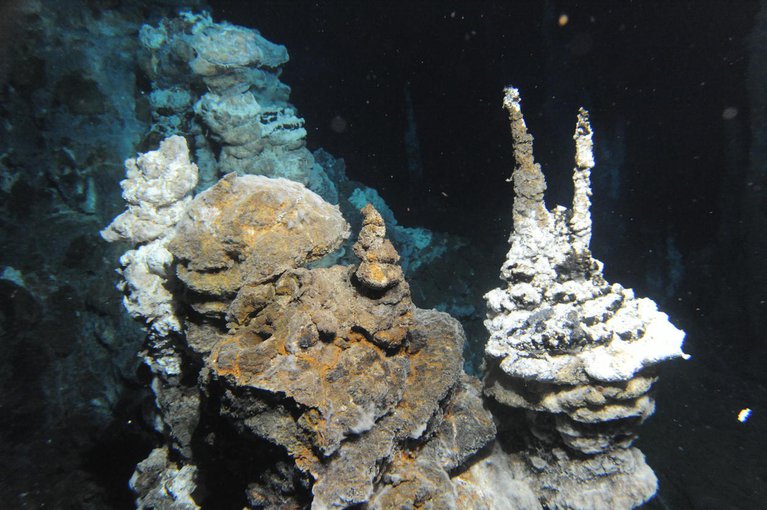

Loki’s Castle.Image credit: Centre for Geobiology by R.B. Pedersen.
A Loki strike
In the end, Ettema and his colleagues were able to assemble a nearly whole genome (92 percent complete) from the bits and pieces of DSAG-targeted DNA. Because the sample came from a site near an active hydrothermal vent called Loki’s Castle, the team decided to call these organisms Lokiarchaea (phylum name: Lokiarchaeota).
In this genome, the researchers identified 5,381 protein coding genes. From these, they selected 36 universal genes (ones found in all life forms), which they used to compare with a set of other organisms from the different three domains of life. To improve the discriminatory power, they pasted the universal-gene codes together into a single “concatenated” set. One can think of this as a bar code that is unique to each organism. If two barcodes are similar, then it means the corresponding organisms are closely related.
The results of this barcode analysis revealed that Lokiarchaeota are highly related to Eukaryota, suggesting we eukaryotes have found a long-lost cousin hiding at the bottom of the ocean.
The researchers performed several consistency checks to convince themselves that what they were seeing was real.
However, Norman Pace, a microbiologist from the University of Colorado, isn’t convinced. He doesn’t think concatenated gene sets can be trusted. “There’s too much random information,” Pace says. “The resulting trees are unreliable.”
Pace believes there’s more certainty in ribosomal DNA. Pace and his co-workers analyzed the 16S rRNA data of Lokiarchaeota that Ettema and colleagues made public. “Our analysis of their data says that the Lokiarchaeota are not something new,” Pace says. They belong to a long-known group of environmental Crenarchaeota.
Phylogenetic trees based on ribosomal RNA genes are more reliable, Pace says, because these genes are one of the most conserved. And ribosomal analyses consistently support a three-domain tree of life. Pace also thinks that three-domains make more sense in explaining why archaea use a form of membrane lipid that is distinct from the lipids used by eukaryotes and bacteria.
Ettema counters that rRNA has its own problems. Past research has identified a high-temperature bias in the 16S rRNA gene, which causes heat-loving organisms to move closer to the base of the tree of life. Moreover, Ettema’s lab has performed follow-up analyses of rRNA gene sequences from Lokiarchaeota and related lineages, and these analyses were found to support the evolutionary relation between Lokiarchaea and eukaryotes.
Eukaryote tool kit?
Ettema also believes he and his colleagues have additional evidence that makes their case. In the Lokiarchaeal genome, the team identified genes that normally only appear in eukaryotes. These genes are traditionally associated with certain complex functions, like cytoskeletal formation and lysosome production.
Ettema makes clear that we don’t know yet what Lokiarchaea do with these genes. Moreover, we can’t assume that the common ancestor of Lokiarchaea and eukaryotes had any eukaryotic traits.
“It is very hard to infer what a gene was doing 2 billion years ago.” Ettema says. He and his co-authors only suggest that Lokiarchaeota’s early ancestors had a rich genomic “starter kit.”
However, molecular biologist Patrick Forterre from the Institute Pasteur in Paris, France, does not think the evidence is strong enough that Lokiarchaea have these eukaryote genes. The trouble, according to Forterre, goes back to metagenomics.
“You’re never sure that you aren’t mixing different organisms,” Forterre says. He claims that possibly no more than 50 percent of the DNA they have is from the actual Lokiarchaeota.
When Forterre and his colleagues looked at individual genes from Lokiarchaeota, without doing concatenation, they found that different genes gave different trees. “It doesn’t make sense that these genes are all from one organism,” Forterre says. These discrepancies suggest that Ettema’s sample was perhaps contaminated with unidentified eukaryotes. Forterre believes this contamination would explain why the sample was found to contain a virus that traditionally only infects eukaryotes.
Ettema says they were extremely careful about this very issue. “We wanted to be 1000 percent sure that we didn’t have contamination from another eukaryote,” he says. They tested for eukaryotic ribosomal markers, but nothing lit up that would suggest a eukaryote was in the sample.
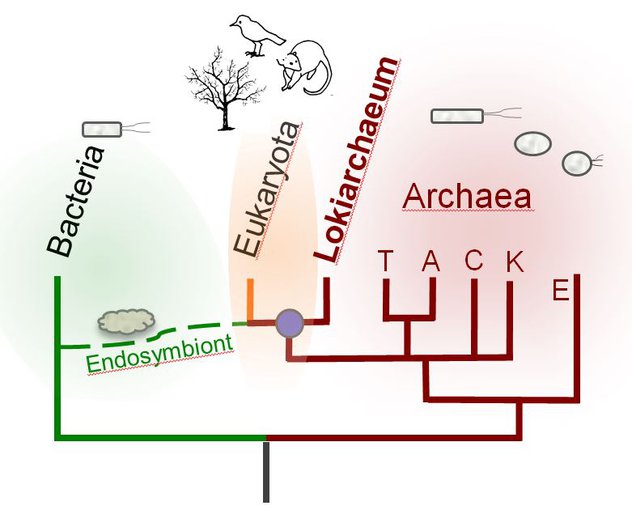
The Lokiarchaea are surprisingly similar to modern eukaryotes, suggesting they share a relatively recent common ancestor. The divergence of Lokiarchaeota and Eukaryota may have coincided with a merger between this common ancestor and a bacteria.Image credit: Christa Schleper, University of Vienna.
Moving forward
As said earlier, naming is an important business in biology. By picking the name “Lokiarchaeota,” Ettema and his colleagues were choosing as namesake the Norse god Loki, which they described in their paper as “a staggeringly complex, confusing, and ambivalent figure who has been the catalyst of countless unresolved scholarly controversies.” The authors apparently foresaw the trouble that the Lokiarchaeota finding would stir up.
But for some scholars, Loki and his controversies are mere tales from the past. “The train has already left the station, and I suspect that the debate over trees is essentially over, except for minor details,” says Lake.
Whether the details are minor or major, the question now is: How to move forward? Whitaker believes there needs to be isolation of a single Lokiarchaeal cell for analysis. “I’d like to see it again,” she says.
Ettema says that is one of their goals now. They are finding Lokiarchaea in other places, like hot springs. He’s hopeful that they will be able to localize one and look at it under a microscope to see if it does anything eukaryote-like, such as engage in dynamic membrane activity.
Whitaker also thinks that we need more data on the other side — from eukaryotes. Lots of genomes are sequenced for macro-organisms, such as primates, farm animals and fruit flies. The unicellular eukaryotes haven’t gotten much attention (unless they are disease-causing).
“We don’t know the diversity of eukaryotes very well,” Whitaker says. “The Tree of Life isn’t complete.”