2015 Annual Science Report
 SETI Institute
Reporting | JAN 2015 – DEC 2015
SETI Institute
Reporting | JAN 2015 – DEC 2015
Rock Sample Biosignature Library and Automated Identification of Biosignatures
Project Summary
The goals of this project are to: 1) build a Raman spectra and imaging library of rock samples containing biosignatures as no publically available online sample library currently exists; and to: 2) use this library as testing and training sets to develop automated classifiers for identifying biosignatures in rock spectra. Building a sample library and developing automated classifiers would enable field scientists or robotic explorers on the surface of Mars or elsewhere to automatically identify biosignatures in rock samples from Raman spectra out in the field.
Project Progress
The development of automated classifiers for identifying biosignatures in rock spectra requires establishing a library of biosignature samples as well as analyzed rock samples containing multiple minerals and biosignatures. Such a custom library is required since no publically available online sample library currently exists. To this end, we have been building such a sample library.
Our sample library currently encompasses over 1200 samples that have undergone careful hand sample analysis and macro imaging under uniform lighting conditions. We are in the process of acquiring Raman spectra of sedimentary samples with our in house dual excitation (532nm and 785nm) Raman instrument provided by EIC Laboratories. Likely, some of these samples will contain biosignatures. Approximately 700 of our samples are igneous rocks, with the remainder being of sedimentary and metamorphic origin. We also have a wide variety of common rock forming minerals that we have analyzed with our Raman instrument. This year, we began to collect a variety of sedimentary samples containing biosignatures, including travertine from Travertine Hot Springs, CA, gypsums and salts from the Atacama Desert containing beta-carotene and other biosignatures, microbial mats, and lab samples of powdered gypsum and anhydrite containing palmitic acid.
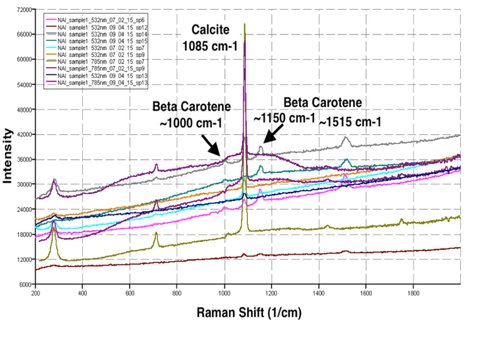
An important component of sample analysis on future missions will be location selection on the sample. Prepared samples are mostly homogeneous, but natural rock samples are more heterogeneous and non-uniform. Analysis of the travertine sample shown in Figure 1 revealed ß-carotene peaks in the more protected interior locations on the rock, while large flatter areas on the outer surface appeared devoid of spectral signatures pointing to life.
As we move away from classifying pure minerals, we need to be able to distinguish individual minerals from biosignatures in the spectra taken from rocks. Therefore, we performed a preliminary test of our existing automated mineral classifier (Ishikawa and Gulick, 2013) to compare them with results from our hand sample analysis. We randomly selected 22 rock samples from our collection and acquired several spectra from each. We tested the ability to detect the presence or absence of plagioclase, potassium feldspar, quartz, and pyroxene in each spectra. This simple test yielded an overall success rate of ~85%. Although we did not have sufficient samples to test for biosignatures, the detection of biosignatures in Raman spectra would likely be similarly successful given enough samples training and testing.
We have also been evaluating numerous other combinations of pre-processing techniques and machine learning approaches (MLA) to improve results especially for biosignature identification, that include cross-validation and voting schemes.
Publications
-
Baker, V. R., Hamilton, C. W., Burr, D. M., Gulick, V. C., Komatsu, G., Luo, W., … Rodriguez, J. A. P. (2015). Fluvial geomorphology on Earth-like planetary surfaces: A review. Geomorphology, 245, 149–182. doi:10.1016/j.geomorph.2015.05.002
-
Rodriguez, J. A. P., Kargel, J. S., Baker, V. R., Gulick, V. C., Berman, D. C., Fairén, A. G., … Glines, N. (2015). Erratum: Martian outflow channels: How did their source aquifers form, and why did they drain so rapidly?. Scientific Reports, 5, 15092. doi:10.1038/srep15092
-
Rodriguez, J. A. P., Kargel, J. S., Baker, V. R., Gulick, V. C., Berman, D. C., Fairén, A. G., … Glines, N. (2015). Martian outflow channels: How did their source aquifers form, and why did they drain so rapidly?. Scientific Reports, 5, 13404. doi:10.1038/srep13404
-
Rodriguez, J. A. P., Platz, T., Gulick, V., Baker, V. R., Fairén, A. G., Kargel, J., … Glines, N. (2015). Did the martian outflow channels mostly form during the Amazonian Period?. Icarus, 257, 387–395. doi:10.1016/j.icarus.2015.04.024
-
PROJECT INVESTIGATORS:
-
PROJECT MEMBERS:
Patrick Freeman
Co-Investigator
Jason Angell
Collaborator
Timothy Johnsen
Collaborator
Paige Morkner
Collaborator
-
RELATED OBJECTIVES:
Objective 1.1
Formation and evolution of habitable planets.
Objective 2.1
Mars exploration.
Objective 7.1
Biosignatures to be sought in Solar System materials